|
The Laboratory for Molecular Modeling (LMM) brings molecular modeling and computational chemistry closer to the experimental scientist. The MM Laboratory uses computer-aided molecular modelling techniques to drive the development of novel bioactive compounds. Research activities focus on the development and the application of modeling techniques in the area of protein-ligand interactions, structure-based drug design, QSAR, homology modeling and bioinformatics. Following their design, candidate molecules are synthesized by a number of laboratories and evaluated in appropriate in vitro and in vivo assays. Advanced modeling software is available to perform quantum calculations on small molecular or solid systems and molecular mechanics/dynamics modeling for large systems such as proteins, DNA, nanomolecules, polymers, solids, and liquids. Additional computing facilities are available and currently used at the Computing Center of CINECA (Casalecchio, Bologna), ISCRA- supercomputing facilities for academic researchers. In addition we are also interest in developing and applying computational methods to examine conformational changes and properties of ligand-binding to the target macromolecular receptor. As an application, in the last few years a particular attention have been directed to application of full atom Molecular dynamics to the study of the conformational and structural characterization of complex supra-macromolecular systems such as membrane receptors (ex. 5-HT2c serotonin receptor). Together with the Bionanotecnology laboratory, another research filed is oriented towards the Design and synthesis of functionalized amphiphilic lipids that can self-assemble into nanostructures such as micelles, liposomes or structures more complex used in gene or drug delivery. Other Facility and research activities of Bionanotecnology Laboratory: Physical chemistry and chemical physics characterization of complex structures and correlation of their structural features with the ability to incorporate drugs or nucleic acids to delivery. Methods used: X-ray diffraction, Electrophoretic and Dynamic Light Scattering, ESR, Electrophoresis, NMR.
|
 |
|
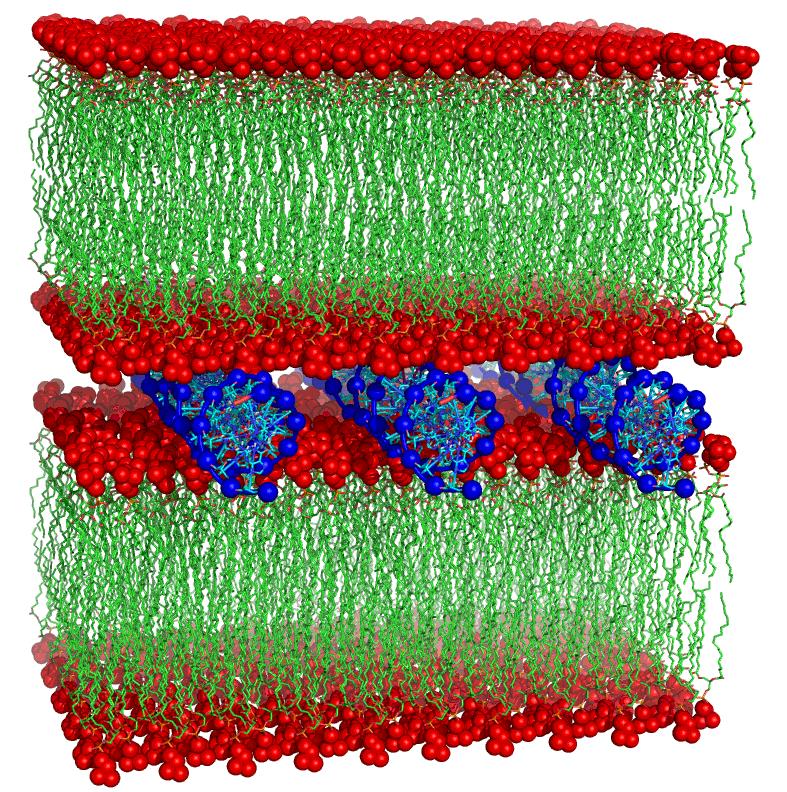
Design and Dynamics strucutral study of mixed composition bilayers as liposomial nanovectors for Gene and Drug Delivery (collaboration with Biotechnology Lab – SIMAU)
• Functionalized liposmes able to complex cations and DNA :
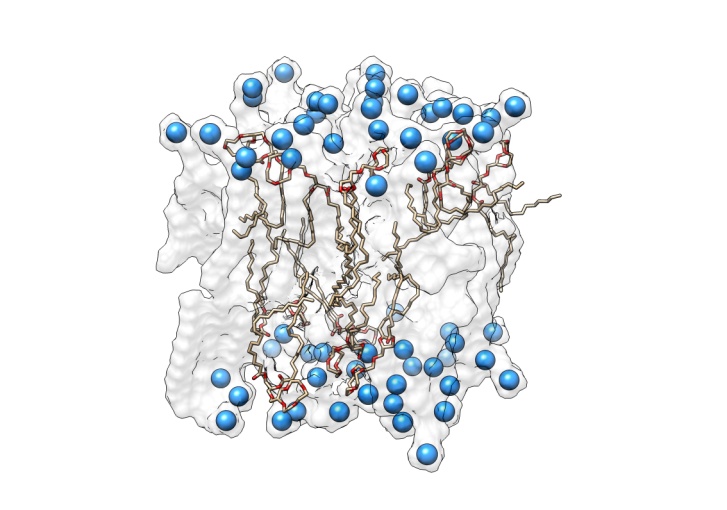
• pH sensible liposomes
• Liposomes conjugated to antioxidants in order to protect against oxidative stress
|
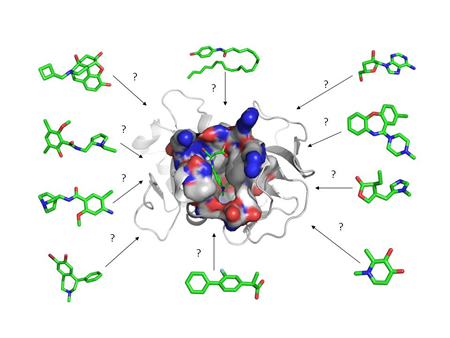
Virtual screening, synthesis and design of new drugs inhibitors of bacterial NaPRT (collaboration with Dr. Leonardo Sorci, Dipartimento di Scienze Cliniche Specialistiche ed Odontostomatologiche and Bionanotecnology lab, SIMAU) |
|
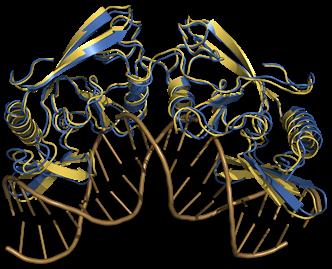
Macromolecular Docking (Protein-Protein, Protein-RNA Docking):
• Development of new protocols for assessing macromolecular association (CAPRI experiments)
• Investigation of the interaction, association mechanism for EIf5 factor and e ribosomial 50S unit in archea
• Investigation of the molecular basis of glutationilation mediated by Human Gliossalase II enzyme.
|
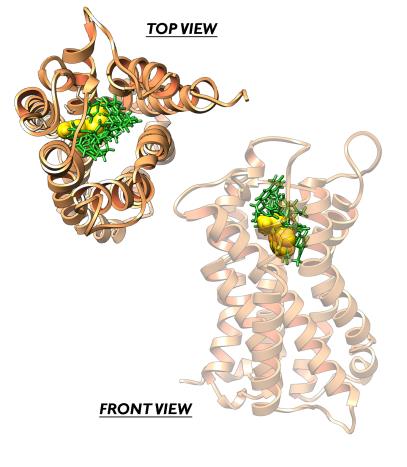
Dynamics of inhibition/activation of GPCRs such as serotonin 5-HT2C
• Binding affinity and influence of genetic polymorphisms in the free energy of association.
• Investigation of the trasdutional of the biological signal mechanism through combined Coarse Grained/full atom MD approach (receptor oligomerization)
|
|
Activity fields (Research Overview)
The research activity that characterizes the Molecular Modelling Laboratory regards the design and development of potential drugs active against enzyme targets of pharmaceutical interest by means of molecular modelling and "de novo" drug design. A distinguishing feature of the Laboratory is the possibility to rationally design "ad hoc" molecules, i.e. molecules designed and developed specifically to interact with a particular macromolecular target. One of the most used approaches is the structure-based design of new molecules by modeling the structures of potential ligands directly into their putative binding site. In many cases, this approach was able to suggest new lead compounds, like enzyme inhibitors, that were further developed as potential drugs through iterative cycles of structure-based optimization. In this respect, an important role is played by docking and virtual screening methods able to predict which molecules are more suited for binding a given target, thereby driving the biological testing process. In the Laboratory, new methodologies have been developed based on combination of different approaches like docking methods, molecular mechanics (MM) and dynamics (MD) methods and more accurate scoring functions, which proved able to screen, for each particular target, many molecules in a reasonable amount of time. In those cases in which the three-dimensional structure of the target is not known experimentally, the Laboratory has the possibility to build 3D models of enzymes and proteins in general through bioinformatics, secondary structure prediction and homology modelling techniques.
In the Laboratory an important role is also given to molecular dynamics simulations, a computational approach important for predicting the three-dimensional structures of macromolecular complexes. These simulations are used to study the structures of target-ligand complexes with the aim of understanding sequence-structure-activity relationships and performing ligand design.
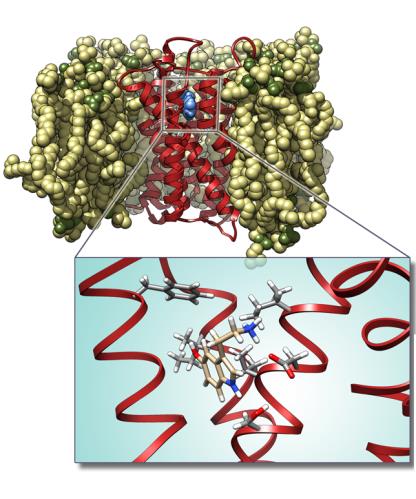
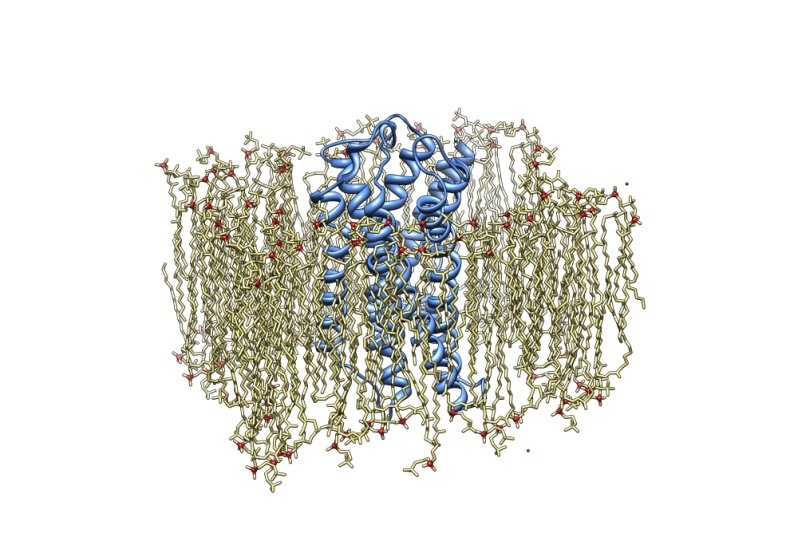
Furthermore, with recent advances in X-ray crystallography of membrane proteins promising many new high-resolution structures, MD simulations become increasingly valuable for understanding membrane protein receptors function. The increase in computational power in synergy with more efficient computational methodologies allows the researcher today to carry out molecular dynamics simulations of any structurally known membrane protein in its native environment i.e. the lipid byilayer, covering the time scale of up to 0.1 μsec. At the frontier of membrane protein simulations are ion channels, G-coupled protein receptors, passive and active transporters, and bioenergetic proteins. In our laboratory, a particular attention have been given to serotonin receptors (particularly 5-HT-2c) and to the membrane protein Aromatase, also involved as target in antitumoral therapy.
Finally, Molecular dynamics simulation of lipid bilayers alone is a rapidly developing field driven by the need to obtain an atomic-level picture of membrane structure and dynamics. As the discipline matures there will be many opportunities to extract information from simulation results that can be utilized in the interpretation and design of laboratory experiments (Bionanotecnology Laboratory, Dr. Pisani Michela, SIMAU, Dr. Giovanna Mobbili, DiSVA). In particular that research is oriented towards the Design and synthesis of functionalized amphiphilic lipids that can self-assemble into nanostructures such as micelles, liposomes or structures more complex used in gene or drug delivery. In all research activities, tight collaborations with research groups working on the experimental validation and development (synthesis, biological activity, crystal structures etc.) have been established.
|